Download this notebook from github.
Basic Usage
Climate indicator computations
xclim is a library of climate indicators that operate on xarray DataArray objects.
xclim provides two layers of computations, one responsible for computations and units handling (the computation layer, the indices), and the other responsible for input health checks and metadata formatting (the CF layer, refering to the Climate and Forecast convention, the indicators). Functions from the computation layer are found in xclim.indices, while indicator objects from the CF layer are found in realm modules (xclim.atmos, xclim.land and xclim.seaIce).
To use xclim in a project, import both xclim and xarray.
[1]:
import xclim
from xclim.testing import open_dataset
import xarray as xr
Indice calculations are performed by opening a netCDF-like file, accessing the variable of interest, and calling the indice function, which returns a new DataArray.
For this example, we’ll first open a demonstration dataset storing surface air temperature and compute the number of growing degree days (the sum of degrees above a certain threshold) at the monthly frequency.
[2]:
# ds = xr.open_dataset("your_file.nc")
ds = open_dataset('ERA5/daily_surface_cancities_1990-1993.nc')
ds.tas
[2]:
<xarray.DataArray 'tas' (location: 5, time: 1461)>
array([[277.49966, 270.44736, 273.5631 , ..., 259.30075, 267.44043, 264.0009 ],
[272.3179 , 268.01813, 273.50452, ..., 249.57759, 258.23706, 260.20535],
[245.21338, 252.72534, 248.18385, ..., 235.18086, 236.17192, 243.2071 ],
[270.79147, 263.67996, 257.4426 , ..., 257.80548, 269.45105, 261.2271 ],
[279.71753, 278.1774 , 279.41824, ..., 280.08725, 280.65396, 280.92868]],
dtype=float32)
Coordinates:
lat (location) float32 44.5 45.5 63.75 52.0 48.5
* location (location) object 'Halifax' 'Montréal' ... 'Saskatoon' 'Victoria'
lon (location) float32 -63.5 -73.5 -68.5 -106.8 -123.2
* time (time) datetime64[ns] 1990-01-01 1990-01-02 ... 1993-12-31
Attributes:
standard_name: air_temperature
long_name: Mean daily surface temperature
units: K
cell_methods: time: mean within days[3]:
gdd = xclim.atmos.growing_degree_days(tas=ds.tas, thresh='10.0 degC', freq='YS')
gdd
[3]:
<xarray.DataArray 'growing_degree_days' (location: 5, time: 4)>
array([[7.9897247e+02, 7.2488672e+02, 6.4941925e+02, 6.7033386e+02],
[1.2330164e+03, 1.3716892e+03, 1.1340271e+03, 1.2288167e+03],
[8.0615845e+00, 2.7421051e+01, 1.0251160e+00, 2.0045013e+01],
[9.3481873e+02, 1.0134860e+03, 7.2482220e+02, 6.3551764e+02],
[6.2461761e+02, 5.3345679e+02, 6.3453369e+02, 5.9410144e+02]],
dtype=float32)
Coordinates:
* time (time) datetime64[ns] 1990-01-01 1991-01-01 1992-01-01 1993-01-01
lat (location) float32 44.5 45.5 63.75 52.0 48.5
* location (location) object 'Halifax' 'Montréal' ... 'Saskatoon' 'Victoria'
lon (location) float32 -63.5 -73.5 -68.5 -106.8 -123.2
Attributes:
units: K days
cell_methods: time: mean within days time: mean within days time: sum o...
history: [2022-01-10 15:45:40] growing_degree_days: GROWING_DEGREE...
standard_name: integral_of_air_temperature_excess_wrt_time
long_name: Growing degree days above 10.0 degc
description: Annual growing degree days above 10.0 degcThis computation was made using the growing_degree_days indicator. The same computation can be made through the indice:
[4]:
gdd = xclim.indices.growing_degree_days(tas=ds.tas, thresh='10.0 degC', freq='YS')
gdd
[4]:
<xarray.DataArray 'tas' (location: 5, time: 4)>
array([[7.9897247e+02, 7.2488672e+02, 6.4941925e+02, 6.7033386e+02],
[1.2330164e+03, 1.3716892e+03, 1.1340271e+03, 1.2288167e+03],
[8.0615845e+00, 2.7421051e+01, 1.0251160e+00, 2.0045013e+01],
[9.3481873e+02, 1.0134860e+03, 7.2482220e+02, 6.3551764e+02],
[6.2461761e+02, 5.3345679e+02, 6.3453369e+02, 5.9410144e+02]],
dtype=float32)
Coordinates:
* time (time) datetime64[ns] 1990-01-01 1991-01-01 1992-01-01 1993-01-01
lat (location) float32 44.5 45.5 63.75 52.0 48.5
* location (location) object 'Halifax' 'Montréal' ... 'Saskatoon' 'Victoria'
lon (location) float32 -63.5 -73.5 -68.5 -106.8 -123.2
Attributes:
units: K dThe call to xclim.indices.growing_degree_days first checked that the input variable units were units of temperature, ran the computation, then set the output’s units to the appropriate unit (here K d or kelvin days). As you can see, the indicator returned the same output, but with more metadata, it also performed more checks as explained below.
The growing_degree_days indice makes most sense with daily input, but could accept other source frequencies. The computational layer assumes that users have checked that the input data has the expected temporal frequency and has no missing values. However, no checks are performed, so the output data could be wrong. If you’re unsure about all those things, a safer bet is to use ``Indicator`` objects from the CF layer, as done in the following section.
New unit handling paradigm in xclim 0.24 for indices
As of xclim 0.24, the paradigm in unit handling has changed slightly. Now, indices are written in order to be more flexible as to the sampling frequency and units of the data. You can use growing_degree_days on, for example, the 6-hourly data. The ouput will then be in degree-hour units (K h). Moreover, all units, even when untouched by the calculation, will be reformatted to a CF-compliant symbol format. This was made to ensure consistency between all indices.
Very few indices will convert their output to a specific units, rather it is the dimensionality that will be consistent. The Unit handling page goes in more details on how unit conversion can easily be done.
This doesn’t apply to Indicators. Those will always output data in a specific unit, the one listed in the Indicators.cf_attrs metadata dictionnary.
Finally, as almost all indices, the function takes a freq argument to specify over what time period it is computed. These are called “Offset Aliases” and are the same as the resampling string arguments. Valid arguments are detailed in panda’s doc (note that aliases involving “business” notions are not supported by xarray and thus could raises issues in xclim.
Health checks and metadata attributes
Indicator instances from the CF layer are found in modules bearing the name of the computational realm in which its input variables are found: xclim.atmos, xclim.land and xclim.seaIce. These objects from the CF layer run sanity checks on the input variables and set output’s metadata according to CF-convention when they apply. Some of the checks involve:
Identifying periods where missing data significantly impacts the calculation and omits calculations for those periods. Those are called “missing methods” and are detailed in section Health checks.
Appending process history and maintaining the historical provenance of file metadata.
Writing Climate and Forecast Convention compliant metadata based on the variables and indices calculated.
Those modules are best used for producing NetCDF that will be shared with users. See Climate Indicators for a list of available indicators.
If we run the growing_degree_days indicator over a non daily dataset, we’ll be warned that the input data is not daily. That is, running xclim.atmos.growing_degree_days(ds.air, thresh='10.0 degC', freq='MS') will fail with a ValidationError:
[5]:
ds6h = xr.tutorial.open_dataset('air_temperature')
gdd = xclim.atmos.growing_degree_days(ds6h.air, thresh='10.0 degC', freq='MS')
---------------------------------------------------------------------------
ValidationError Traceback (most recent call last)
/tmp/ipykernel_650/3535010228.py in <module>
1 ds6h = xr.tutorial.open_dataset('air_temperature')
----> 2 gdd = xclim.atmos.growing_degree_days(ds6h.air, thresh='10.0 degC', freq='MS')
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/indicator.py in __call__(self, *args, **kwds)
645
646 # Pre-computation validation checks on DataArray arguments
--> 647 self._bind_call(self.datacheck, **das)
648 self._bind_call(self.cfcheck, **das)
649
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/indicator.py in _bind_call(self, func, **das)
776 else:
777 # Call the func using bound arguments
--> 778 return func(*ba.args, **ba.kwargs)
779
780 @classmethod
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/indicator.py in datacheck(**das)
1085 for key, da in das.items():
1086 if "time" in da.coords and da.time.ndim == 1 and len(da.time) > 3:
-> 1087 datachecks.check_daily(da)
1088
1089
<boltons.funcutils.FunctionBuilder-142> in check_daily(var)
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/options.py in run_check(*args, **kwargs)
114 @wraps(func)
115 def run_check(*args, **kwargs):
--> 116 return _run_check(func, DATA_VALIDATION, *args, **kwargs)
117
118 return run_check
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/options.py in _run_check(func, option, *args, **kwargs)
106 func(*args, **kwargs)
107 except ValidationError as err:
--> 108 raise_warn_or_log(err, OPTIONS[option], stacklevel=4)
109
110
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/utils.py in raise_warn_or_log(err, mode, msg, stacklevel)
267 warnings.warn(msg, stacklevel=stacklevel + 1)
268 else: # mode == "raise"
--> 269 raise err
270
271
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/options.py in _run_check(func, option, *args, **kwargs)
104 """Run function and customize exception handling based on option."""
105 try:
--> 106 func(*args, **kwargs)
107 except ValidationError as err:
108 raise_warn_or_log(err, OPTIONS[option], stacklevel=4)
~/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/datachecks.py in check_daily(var)
43 """
44 if xr.infer_freq(var.time) != "D":
---> 45 raise ValidationError("time series is not recognized as daily.")
ValidationError: time series is not recognized as daily.
Resampling to a daily frequency and running the same indicator succeeds, but we still get warnings from the CF metadata checks.
[6]:
daily_ds = ds6h.resample(time='D').mean(keep_attrs=True)
gdd = xclim.atmos.growing_degree_days(daily_ds.air, thresh='10.0 degC', freq='YS')
/home/docs/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/cfchecks.py:40: UserWarning: Variable does not have a `cell_methods` attribute.
vardata, "cell_methods", parse_cell_methods(data["cell_methods"]) + "*"
/home/docs/checkouts/readthedocs.org/user_builds/xclim/envs/v0.29.0/lib/python3.7/site-packages/xclim/core/cfchecks.py:43: UserWarning: Variable does not have a `standard_name` attribute.
check_valid(vardata, "standard_name", data["standard_name"])
To suppress the CF validation warnings in the following, we will set xclim to send them to the log, instead of raising a warning or an error.
The missing method which determines if a period should be considered missing or not can be controlled through the check_missing option, globally or contextually. The main missing methods also have options that can be modified.
[7]:
with xclim.set_options(check_missing='pct', missing_options={'pct': dict(tolerance=0.1)}, cf_compliance='log'):
# Change the missing method to "percent", instead of the default "any"
# Set the tolerance to 10%, periods with more than 10% of missing data
# in the input will be masked in the ouput.
gdd = xclim.atmos.growing_degree_days(daily_ds.air, thresh='10.0 degC', freq='MS')
Finally, xclim also allows to call indicators using datasets and variable names.
[8]:
with xclim.set_options(cf_compliance='log'):
gdd = xclim.atmos.growing_degree_days(tas='air', thresh='10.0 degC', freq='MS', ds=daily_ds)
# variable names default to xclim names, so we can even do this:
renamed_daily_ds = daily_ds.rename(air='tas')
gdd = xclim.atmos.growing_degree_days(thresh='10.0 degC', freq='MS', ds=renamed_daily_ds)
Graphics
[9]:
import matplotlib.pyplot
%matplotlib inline
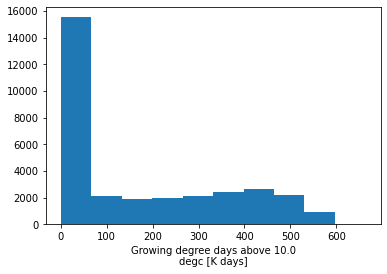
# Summary statistics histogram
gdd.plot()
[9]:
(array([15532., 2079., 1861., 1935., 2137., 2416., 2675., 2202.,
931., 32.]),
array([ 0. , 66.32573, 132.65146, 198.9772 , 265.30292, 331.62866,
397.9544 , 464.28012, 530.60583, 596.9316 , 663.2573 ],
dtype=float32),
<BarContainer object of 10 artists>)
[10]:
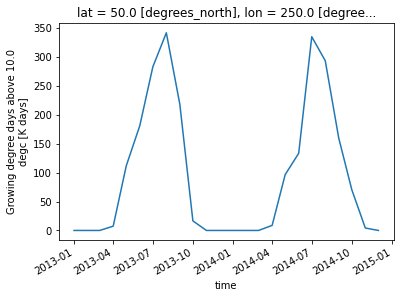
# Show time series at a given geographical coordinate
gdd.isel(lon=20, lat=10).plot()
[10]:
[<matplotlib.lines.Line2D at 0x7f10232c2e10>]
[11]:
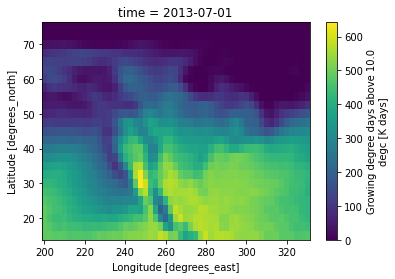
# Show spatial pattern at a specific time period
gdd.sel(time='2013-07').plot()
[11]:
<matplotlib.collections.QuadMesh at 0x7f10235c9290>
For more examples, see the directions suggested by xarray’s plotting documentation
To save the data as a new NetCDF, use to_netcdf.
[12]:
gdd.to_netcdf('monthly_growing_degree_days_data.nc')
It’s possible to save Dataset objects to other file formats. For more information see: xarray’s documentation